A study accepted to ML4H 2025 explores whether sequence-based models trained on large-scale T-cell activation data can improve prediction of T cell receptor recognition, a longstanding challenge in computational immunology with implications for cancer and autoimmune disease.
Predicting how T cell receptors (TCRs) recognize peptide–MHC (pMHC) antigens is one of the central challenges in immunology today. It is also one of the hardest to solve. T cell recognition arises from complex molecular interactions, and even recent advances in protein structure prediction have not fully resolved which receptor-antigen pairs will lead to T cell activation. That question has broad relevance across immune-mediated diseases. In cancer, it could help researchers better understand antigen-specific immune responses relevant to target discovery and therapeutic design, while in autoimmune disease it may offer new ways to study the recognition events that drive immune dysregulation.
A recent Adaptive paper presented at Machine Learning for Health (ML4H) Symposium 2025 examines that challenge using ImmSET (Immune Synapse Encoding Transformer), a sequence-based transformer model trained on large-scale T-cell activation data. The research asks what becomes possible when models are trained directly on large-scale T-cell datasets rather than relying primarily on inferred structure.
What did the study examine?
Recent AI tools have expanded what is possible in structural biology. Deep learning systems such as AlphaFold have improved researchers’ ability to model proteins and protein complexes, opening new avenues for studying molecular function. But T cell recognition remains difficult to predict because the problem is not simply one of structure. It also requires understanding which receptor-antigen interactions are associated with T cell activation.
ImmSET was designed to test whether a sequence-based model could learn that relationship from data at scale. Trained from scratch using one of Adaptive’s large proprietary T-cell activation datasets, the model uses sequence information alone to predict T-cell receptor recognition of pMHC antigens.
Three findings from the study are especially notable.
Finding 1: Sequence-based prediction outperformed structure-based pipelines on one benchmark
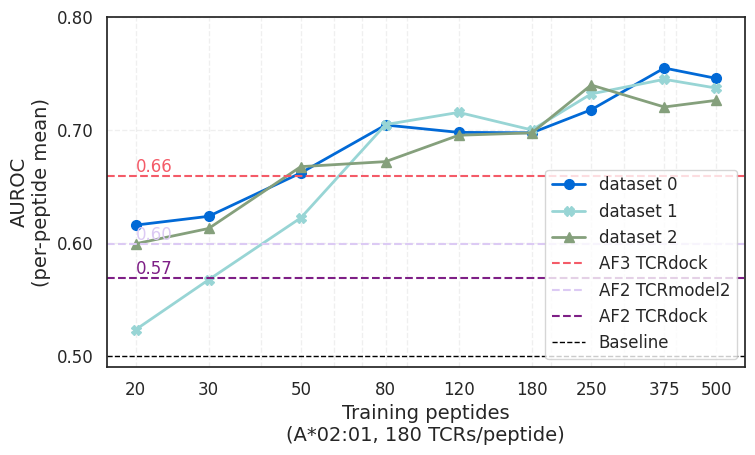
When provided a sufficiently large training dataset, ImmSET outperformed AlphaFold2- and AlphaFold3-based pipelines on a benchmark assessing HLA-A*02:01 specificity prediction, while running orders of magnitude faster.
That result is notable because structure-based approaches have come to represent a major frontier in biological prediction. The study suggests that, for this task, sequence-based models trained on large T-cell activation datasets can provide a practical and scalable alternative. Rather than replacing structural approaches, the findings point to a complementary path—one that may be especially useful where speed, throughput, and dataset scale matter.
Finding 2: Performance improved as the dataset grew
A second finding was the consistency with which model performance improved as training data increased. The study reports performance gains as both the number of peptides increased and as the number of TCRs observed per peptide increased.
This suggests the model is not nearing an early ceiling. Instead, it points to a clear path for continued improvement as larger and more diverse T-cell activation datasets become available. In a field where data limitations have often constrained progress, that is an important signal.
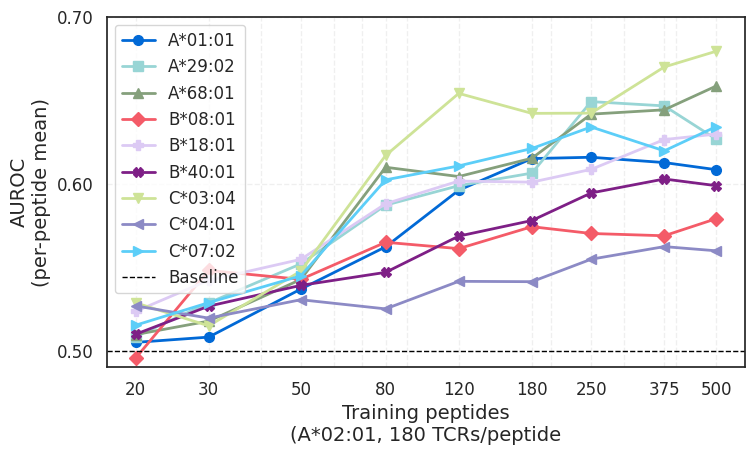
Finding 3: Predictive signal began to extend across HLA alleles
The study also found evidence that increasing scale improved generalization beyond the allele on which the model was trained. Although ImmSET was trained only on A*02:01 data, the authors report measurable predictive signal on other HLA alleles as training data increased.
The cross-allele signal remains early, but it suggests the model may be learning features of T cell recognition that are generalizable across HLA alleles. HLA varies widely across the human population, so models need to do more than learn from the specific alleles in their training data. They need to generalize beyond them.
Why evaluation matters
The paper also addresses a methodological issue that has complicated prior work in this area: shortcut learning. According to the authors, some sequence-based models can appear to generalize even when peptide information is removed, effectively relying on memorized TCR motifs rather than learning meaningful receptor-antigen relationships.
To address that risk, the study emphasizes strict evaluation on truly unseen peptides. That is an important point. In problems as complex as TCR recognition, apparent performance gains can be misleading if evaluation is not designed carefully enough to test real generalization.
Why this matters across cancer and autoimmune disease
Taken together, the findings point to a practical direction for the field. Structural models have opened an important chapter in computational biology, but predicting immune recognition may also require approaches trained directly on large-scale T-cell activation data. The paper does not suggest that the mechanistic complexity of T cell recognition has been solved. It does suggest, however, that sequence-based models can provide fast, scalable, and increasingly generalizable predictions that complement structure-based methods as the field evolves.
That direction matters across immunology. In cancer, better prediction of T cell recognition could help inform antigen selection, including for personalized cancer vaccines, as well as T cell selection for cell therapies. In autoimmune disease, the same challenge applies in a different context: understanding which recognition events may contribute to pathogenic immune responses. Across both, improved prediction could help researchers study immune recognition more systematically.
Accepted and presented at ML4H 2025, the study adds to a growing body of work suggesting that progress in computational immunology may depend not only on better models, but on larger and more informative datasets.
Read the full paper on OpenReview.
