Harnessing the power of immune receptor data
Our AI-enabled immune medicine platform offers high-throughput deep sequencing of T-cell and B-cell receptors at unparalleled scale to enable comprehensive immune profiling, with broad applications in oncology, virology, and autoimmune research.
The adaptive immune system is massively diverse
The adaptive immune system is incredibly diverse, shaped by a lifetime of exposures and responses to different antigens. Unlike conventional immune assays that capture only a fraction of that diversity, Adaptive’s immune receptor sequencing technologies unlock the ability to accurately measure, track and detect patterns of an individual’s repertoire dynamics.
Your end-to-end immune medicine partner
Adaptive’s immune medicine platform empowers research at all stages from preclinical development through Phase 3 clinical trials. Our T-cell and B-cell receptor sequencing assays profile the adaptive immune repertoire with unparalleled scale and precision, delivering rich insights that enable data-driven decisions in oncology, infectious disease, autoimmune conditions, and more.
Broad clinical applications
Investigate mechanisms of action and biomarkers of response.
Bias-controlled immune profiling assays
High-throughput sequencing for reliable and reproducible results.
Guided data analysis & interpretation
Expert computational biology support to maximize your results.
Our core services
Adaptive’s Immune Medicine platform covers the full research and clinical pipeline – from preclinical target discovery to translational diagnostics.
T-cell & B-cell receptor deep sequencing
Our robust T- and B-cell sequencing assays accurately measure the breadth and depth of an individual’s adaptive immune response. This immune profiling data informs broad applications in cancer, infectious diseases and autoimmunity.
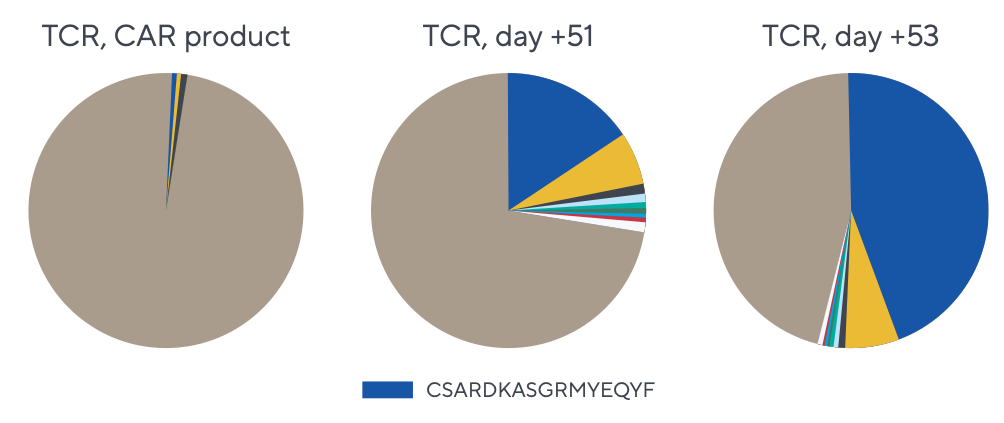
Repertoire data analysis
With more than a decade of deep domain expertise, Adaptive helps partners turn rich, comprehensive immune receptor data into biologically meaningful insights that support actionable decisions. Explore two examples of these analyses below.
TCR-antigen data annotation at scale
Using our AI-enabled immune medicine platform, Adaptive has amassed one of the largest curated TCR datasets that maps to thousands of disease antigens in cancer, virology and autoimmune conditions.
Target discovery in autoimmunity
Adaptive has successfully identified the likely disease-causing, ‘autoreactive’ TCRs and the antigens they bind in several autoimmune conditions, such as spondyloarthritis, type 1 diabetes and multiple sclerosis, among others.
The most published immune medicine platform in the world
Peer-reviewed publications and counting
Oncology
Immunology
Autoimmunity
Infectious disease
Published in the world's leading peer-reviewed journals

Immune repertoire data that drives R&D decisions
Your immune receptor sequencing data is fully yours—secure, accessible, and ready to explore. Connector is our streamlined portal for direct access to repertoire data and secure, two-way file sharing. Visualizer enables users to interactively explore their data, generate downloadable figures, and uncover new insights, making immune repertoire analysis accessible to every team.
Translational science insights from immune repertoire data
Interpreting immune profiling data to deliver meaningful translational insights.
Ready to advance your immune research?
Connect with our experts and tell us about your research needs.
Request A Consultation
Frequently Asked Questions
What is immune receptor sequencing?
Immune receptor sequencing is an immune profiling method that uses high-throughput sequencing to characterize the B-cell and T-cell repertoire at scale. Adaptive’s T- and B-cell receptor sequencing assays target the complementary determining region 3 (CDR3), the most diverse part of an immune receptor that determines antigen specificity and serves as a unique bar code to track clonal lineages.
How do gDNA- and RNA-based sequencing methods compare?
With gDNA as the input, Adaptive’s bias-controlled assay can provide absolute cell counts, enabling accurate assessment of immune cell density, repertoire diversity, and clonal expansion. Complementary DNA or RNA-based measurements are obscured by cell expression, which can confound some analyses of repertoire diversity or clonal dynamics.
Can you identify what antigens T-cells are responding to?
Using both antigen stimulation experimental data and computational machine learning approaches, Adaptive has curated an extensive immunomics database that maps T-cell receptor sequences with their corresponding antigen. Your sequenced T-cell repertoires can be queried against this database to elucidate associations with the mapped exposures. For more information, refer to our T-cell receptor repertoire annotations document.
What sample types are compatible with your sequencing assays?
Adaptive’s T- and B-cell receptor sequencing assays are compatible with a wide range of fresh and archival specimen types, including whole blood, peripheral blood mononuclear cells, tissue (frozen, FFPE, or OCT), sorted cells, and more. Refer to our assay-specific guidelines below for more information.
Human TCRB sample guidelines
Human IGH sample guidelines
Mouse TCRB sample guidelines
How do I send my samples to Adaptive?
After you have initiated the order process with our team, you can ship your samples to Adaptive using the courier of your choice. Except for U.S. federal holidays, Adaptive can receive samples Monday–Friday 9am-5pm or Saturday 9am–12pm. We recommend sending samples via overnight or express service to arrive within these hours. For more information, see our Sample packing and shipping instructions.
What data analysis support does Adaptive provide?
The Translational Data Sciences team at Adaptive works directly with our partners to analyze and interpret their immune receptor sequencing data. These analyses include quantifying repertoire diversity, tracking longitudinal trends, and highlighting cohort differences. Incorporating information such as treatment arms or patient outcomes can translate these findings into clinically relevant, actionable insights.